What are Protein Interaction Networks?Proteins interaction networks refer to complex systems of proteins which interact with one another, either directly or indirectly, in order to perform a variety of cellular functions. These interaction networks serve useful to researchers in identifying new functions for proteins based on the known functions of proteins which they interact with.
How do you look at Protein Interaction Networks?Protein interaction networks can be examined using the site STRING, which takes a protein of interest, and creates protein networks based on documented and predicted protein-protein interactions (Figure 1). STRING is linked with the gene ontology database, and through a quick look in the analysis section, different biological processes can be selected, which results in a highlight over all linked genes which may participate in the biological process. By hovering and clicking on certain proteins within the network, each proteins known functions will appear within a short synopsis, further elucidating on related protein functions.
|
Figure 1: Protein Interaction Network of GFAP produced using STRING [3].
|
|
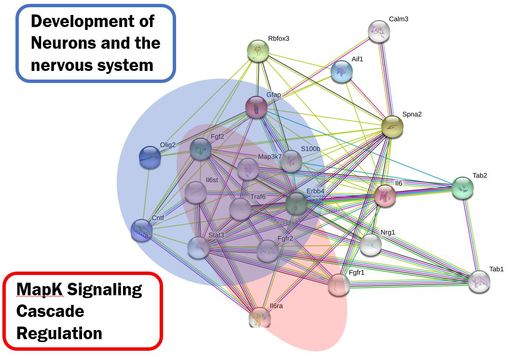
Figure 2: Highlighted Gene Ontology processes in a Human GFAP protein interaction network generated by STRING.
DiscussionDue to the conserved nature of GFAP among humans and mice, it is unsurprising they share many of the same proteins in their interaction networks that participate in similar biological processes. Looking at both networks, GFAP appears to participate in a wide variety of interactions involving neuronal and nervous system development. This makes sense given GFAPs primary expression within the brain and the nervous system. One interesting thing that is different between the networks is the lack of prion protein interactions with GFAP in the mouse STRING network, providing another interaction of interest to test in mice.
|
Which proteins does GFAP interact with?Using STRING, Human and mouse GFAP protein interaction networks were created and analyzed according to their gene ontology terms. Proteins involved in neuronal development and nervous system development were found in both human and mouse string networks, as well as proteins which are part of the Fc-epsilon/MapK signaling pathways (Figures 2 and 3). Most of these proteins were not only part of the same signaling pathways, but also had homologs that were found in both of the string networks.
Figure 3: Highlighted Gene Ontology processes in a Mouse GFAP protein interaction network generated by STRING.
|
References
[1] Gene Ontology Consortium. (n.d.). Retrieved April 12, 2018, from http://geneontology.org/
[2] Skop, A. (2018). Lab 7: Protein Interaction Networks. Retrieved April 11, 2018, from http://genetics564.weebly.com/interaction-networks.html
[3] STRING. (2018, April 11). Retrieved April 11, 2018, from http://string-db.org/
[2] Skop, A. (2018). Lab 7: Protein Interaction Networks. Retrieved April 11, 2018, from http://genetics564.weebly.com/interaction-networks.html
[3] STRING. (2018, April 11). Retrieved April 11, 2018, from http://string-db.org/