What is Homology?
|
Homology is an area of study and a scientific concept that looks at the similarities of genes, proteins, and structures across different species, and infers common evolutionary ancestry and sometimes shared function [2]. Gene, proteins, and structures of different species which share an unusually high percentage of similarity are homologous, and they are homologs of one another. The study of shared function between homologs, is especially relevant in the case of genes and proteins when trying to pick specific model organisms. Below is a list of protein homologs of GFAP which includes a few common model organisms.
|
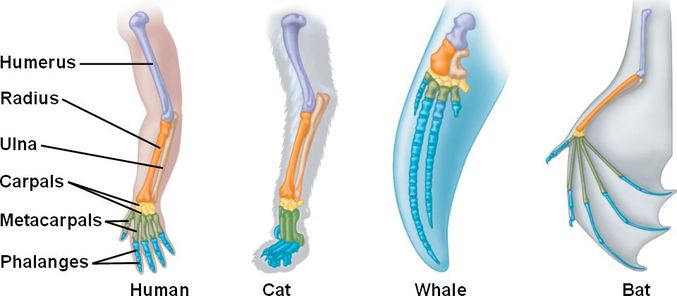
Figure 1: Comparison of different species forelimbs shows structural homology.
|
Protein Homologs of GFAP
|
Drosophilia melanogaster (Fruit Fly)
lamin C, isoform A Accession: NP 523742.2 Length: 621aa Identity: 31% Mus musculus (House Mouse)
glial fibrillary acidic protein isoform 2 Accession: NP_034407.2 Length: 430aa Identity: 91% Rattus norvegicus (Brown rat)
glial fibrillary acidic protein Accession: NP_058705.2 Length: 430aa Identity: 92% Latimeria chalumnae (Coelacanth)
glial fibrillary acidic protiein Accession: XP_006000386.1 Length: 359aa Identity: 73% Cricetulus griseus (Chinese Hamster)
glial fibrillary acidic protein isoform X2 Accession: XP_003504558.1 Length: 430aa Identity: 92% |
Homo sapiens (Human)
glial fibrillary acidic protein isoform 1 Accession: NP_002046.1 Length: 432aa Identity: 100% Pan troglodytes (Chimpanzee)
glial fibrillary acidic protein isoform X1 Accession: XP_511561.4 Length: 432aa Identity: 100% Danio rerio (Zebrafish)
glial fibrillary acidic protein Accession: AAN87836.1 Length: 444aa Identity: 66% Macaca mulatta (Rhesus Monkey)
glial fibrillary acidic protein isoform X1 Accession: XP_014975378.1 Length: 502aa Identity: 99% Felis catus (Domestic Cat)
glial fibrillary acidic protein isoform X2 Accession: XP_003997044.1 Length: 432aa Identity: 95% Loxodonta africana (Elephant)
glial fibrillary acidic protein isoform X2 Accession: XP_003414340.1 Length: 432aa Identity: 96% |
Gallus gallus (Chicken)
glial fibrillary acidic protein Accession: XP_418091 Length: 422aa Identity: 70% Equus caballus (Horse)
glial fibrillary acidic protein Accession: NP_001157333.1 Length: 427aa Identity: 95% Caenorhabditis elegans (Roundworm)
intermediate filament protein 1 Accession: NP 741903.1 Length: 575aa Identity: 33% Ailuropoda melanoleuca (Panda)
glial fibrillary acidic protein isoform X1 Accession: XP_002919795.1 Length: 432aa Identity: 95% |
What is BLAST?BLAST stands for Basic Local Alignment Search Tool, and it is an algorithm commonly used in bioinformatics, an area of research which combines biological information with technological analysis. BLAST takes biological sequence information of a gene or protein of interest, and then compares the sequence information against a selected organisms genome and proteome [1]. This comparison produces a BLAST alignment, which is used to determine possible homologs of your gene of interest that are found in the selected organism's sequence information. Once the potential homolog with the highest percent similarity is identified, a reverse BLAST must be performed. In a reverse BLAST the potential homolog's sequence is compared against the organism where the gene or protein of interest was originally found. If the first result in the reverse BLAST alignment is the gene of interest, then these two sequences are considered homologs of one another [1].
|
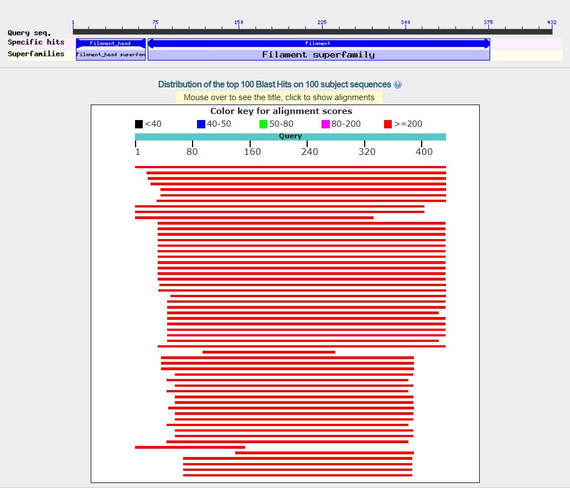
Figure 2: Basic Local Alignment Search Tool (BLAST) alignment
of potential GFAP homologs in mice. |
Discussion of Protein Homology
GFAP shows a high conservation among a wide variety of different vertebrate organisms, but no true homologs of GFAP were found in invertebrates including the commonly used model organisms, the fruit fly and C. elegans. The high conservation of homologs of GFAP in only vertebrates, along with its specific expression localized to the brain, indicates it may play a specialized role in the proper central nervous system development of vertebrates.
References
[1] Basic Local Alignment Search Tool. (n.d.). Retrieved March 14, 2018, from http://blast.ncbi.nlm.nih.gov/Blast.cgi
[2] Pearson, W. R. (2013). An Introduction to Sequence Similarity (“Homology”) Searching. Current Protocols in Bioinformatics / Editoral Board, Andreas D. Baxevanis ... [et Al.], 0 3. https://doi.org/10.1002/0471250953.bi0301s42
[3] Skop, A. (2018). Lab 2: Homology & Phylogeny. Retrieved March 14, 2018, from http://genetics564.weebly.com/homology--phylogeny.html
[2] Pearson, W. R. (2013). An Introduction to Sequence Similarity (“Homology”) Searching. Current Protocols in Bioinformatics / Editoral Board, Andreas D. Baxevanis ... [et Al.], 0 3. https://doi.org/10.1002/0471250953.bi0301s42
[3] Skop, A. (2018). Lab 2: Homology & Phylogeny. Retrieved March 14, 2018, from http://genetics564.weebly.com/homology--phylogeny.html