What are protein domains?
Protein domains are specialized structural units that make up a proteins functional components [1,2]. These domains are usually responsible for a particular job or interaction the protein participates in, and they define the overall role of each specific protein. Domains are interesting due to the fact similar domains may be found on proteins with vastly different functions [1,2].
Domain structure of GFAP
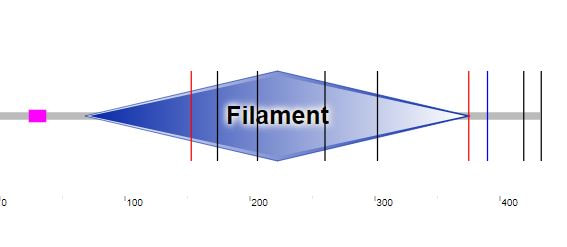
Figure 1: GFAP domain derived from SMART.
Domains for GFAP were found by placing the protein sequence into both SMART and Pfam, bioinformatic sites which are utilized to identify protein domains in a protein of interest [3,4]. GFAP was found to have two domains in both SMART and Pfam, but SMART was only able to define the Filament domain by name and it listed the Filament Head domain as a possible domain. Pfam found both the Filament and the Filament Head domain that comprised the two domains of GFAP.
Figure 2: GFAP domain derived from Pfam.
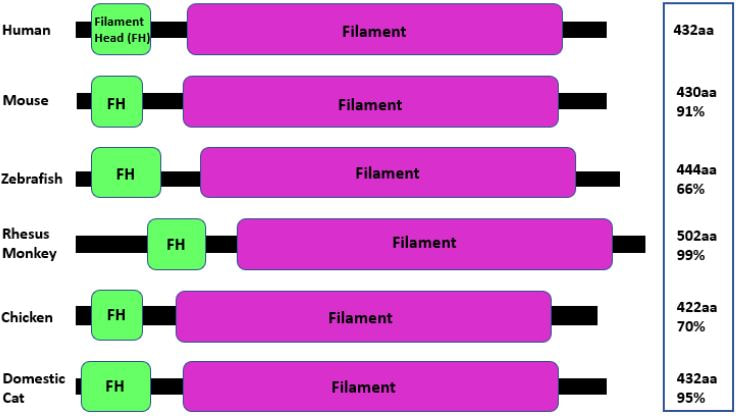
Is GFAP conserved among different species?Looking at the protein domains generated using data from SMART and Pfam a domain alignment was generated. The domain alignment includes percent similarity and the length of each protein. Comparing the domains it is obvious that the Filament Head and Filament domain are both conserved among both the mouse and zebrafish model organisms, as well as in the three additional vertebrates.
|
Figure 2: Conserved domains of GFAP with protein length and percent similarity created using information from BLAST.
|
Discussion
Knowing the specific domains which make up a particular protein can denote the proteins possible functions. GFAP contains two domains common among intermediate filament proteins, which is expected as GFAP is a brain specific intermediate filament protein. The highly conserved nature of GFAP, along with the knowledge of terms derived from Gene Ontology provides insight into the many biological functions of the brain these domains must play a role in. This is likely the reason for why mutations in the domains of GFAP result in Alexander Disease.
References
[1] Sangrador, A. (n.d.). Protein classification: An introduction to EMBL-EBI resources. Retrieved April 12, 2016, from http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi
[2] Skop, A. (2018).Lab 3: Protein domain discovery & DNA motifs. Retrieved April 12, 2016, from http://genetics564.weebly.com/motif--domain-discovery.html
[3] The Pfam protein families database: towards a more sustainable future: R.D. Finn, P. Coggill, R.Y. Eberhardt, S.R. Eddy, J. Mistry, A.L. Mitchell, S.C. Potter, M. Punta, M. Qureshi, A. Sangrador-Vegas, G.A. Salazar, J. Tate, A. Bateman Nucleic Acids Research (2017) Database Issue 44:D279-D285, from http://pfam.xfam.org/
[4] Select your default SMART mode. (n.d.). Retrieved April 12, 2016, from http://smart.embl-heidelberg.de/
[2] Skop, A. (2018).Lab 3: Protein domain discovery & DNA motifs. Retrieved April 12, 2016, from http://genetics564.weebly.com/motif--domain-discovery.html
[3] The Pfam protein families database: towards a more sustainable future: R.D. Finn, P. Coggill, R.Y. Eberhardt, S.R. Eddy, J. Mistry, A.L. Mitchell, S.C. Potter, M. Punta, M. Qureshi, A. Sangrador-Vegas, G.A. Salazar, J. Tate, A. Bateman Nucleic Acids Research (2017) Database Issue 44:D279-D285, from http://pfam.xfam.org/
[4] Select your default SMART mode. (n.d.). Retrieved April 12, 2016, from http://smart.embl-heidelberg.de/